General research interests
My research focuses on the evolution of community-wide interaction architecture and complex, emergent community function across scales of biological organization. From cellular regulation to the ecology of microbiota, interactions are mounted in response to noisy signals and exhibit remarkable context-dependence and historical contingency. These aspects are particularly important in our biomedical microbiome research.
Explaining how complex functions of biological communities emerge from the interactions among their parts is a fundamental question in systems biology and my principal focus. Therefore, I link conceptual modeling with data-driven approaches to gain predictive understanding and to generate testable hypotheses. We integrate concepts from a range of disciplines including microbiology, immunology, ecology, evo-devo, computer science, applied mathematics and physics to better dissect the mechanics of communities and inspire cross-disciplinary knowledge exchange.
Keywords:
interactions | biological complexity | microbial communities | gut-lung axis | systems dynamics | regulatory motifs | multi-omics | network science | mathematical modeling
Reduction of systemic inflammation by elexacaftor/tezacaftor/ivacaftor correlates with lung function improvement in cystic fibrosis.
Halle O, Graeber SY, Kontsendorn J, Kessemeier C, Falke JN, Schwabe J, Schütz K, Pallenberg ST, Dalferth R, Grychtol R, Ringshausen FC, Stahl M, Thee S, Roehmel JF, Syunyaeva Z, Duerr J, Chung J, Hirtz S, Uselmann T, Kühbandner I, Rückes-Nilges C, Bagheri-Pothoff A, Barth S, Schaub B, Brinkmann F, Weber S, Koningsbruggen-Rietschel SV, Abdo M, Weckmann M, Widder S, Hansen G, Tümmler B, Sommerburg O, Naehrlich L, Mall MA, Dittrich AM. Eur Respir J. 2025 Sep 18:2500150. doi: 10.1183/13993003.00150-2025.
Chronic skin and systemic inflammation modulated by S100A8 and S100A9 complexes.
Palomo-Irigoyen M, Bakiri L, Hendrikx T, Hayer S, Schaffenrath J, Widder S, Bachg S, Simmons J, Gallo RL, Starkl P, Roth J, Wagner EF. Cell Death Differ. 2025 Apr 11. doi: 10.1038/s41418-025-01504-9
Microbial community organization designates distinct pulmonary exacerbation types and predicts treatment outcome in cystic fibrosis.
Widder S, Carmody LA, Opron K, Kalikin LM, Caverly LJ, LiPuma JJ. Nat Commun. 2024 Jun 7;15(1):4889. doi: 10.1038/s41467-024-49150-y.
Microbial community organization designates distinct pulmonary exacerbation types and predicts treatment outcome in cystic fibrosis.
Widder S, Carmody LA, Opron K, Kalikin LM, Caverly LJ, LiPuma JJ. Nature Communications. 2024
Longitudinal microbial and molecular dynamics in the cystic fibrosis lung after Elexacaftor-Tezacaftor-Ivacaftor therapy.
Martin C, Guzior DV, Gonzalez CT, Okros M, Mielke J, Padillo L, Querido G, Gil M, Thomas R, McClelland M, Conrad D, Widder S, Quinn RA. Respiratory Research. 2023
Forecasting the dynamics of a complex microbial community using integrated meta-omics.
Delogu F, Kunath BJ, Queirós PM, Halder R, Lebrun LA, Pope PB, May P, Widder S, Muller EEL, Wilmes P. Nature Ecology Evolution. 2024
Metagenomic sequencing reveals time, host, and body compartment-specific viral dynamics after lung transplantation. S Widder, I Goerzer, B Friedel et al. and E. Puchammer-Stoeckl Microbiome 2022
Association of bacterial community types, functional microbial processes and lung disease in cystic fibrosis airways. S Widder, J Zhao, L Carmody, et al. and J J LiPuma. ISMEJ 2021
Multi-omics profiling predicts allograft function after lung transplantations. M Watzenboeck, A Gorki, F Quattrone, R Gawish et al. and S Widder & S Knapp. European Respiratory Journal 2021
Editorial: Microbial Metabolites in Cystic Fibrosis: A Target for Future Therapy? S Widder and S Knapp. American Journal of Respiratory Cell and Molecular Biology 2019
Signatures of ecological processes in microbial community time series. K Faust, F Bauchinger, B Laroche, S de Buyl, L Lahti, A Washburne, D Gonze and S Widder. Microbiome 2018
Using metabolic networks to resolve ecological properties of microbiomes. E Muller, K Faust, S Widder, M Herold, S Martinez Abas and P Wilmes. Cur Opin Sys Biol. 2017
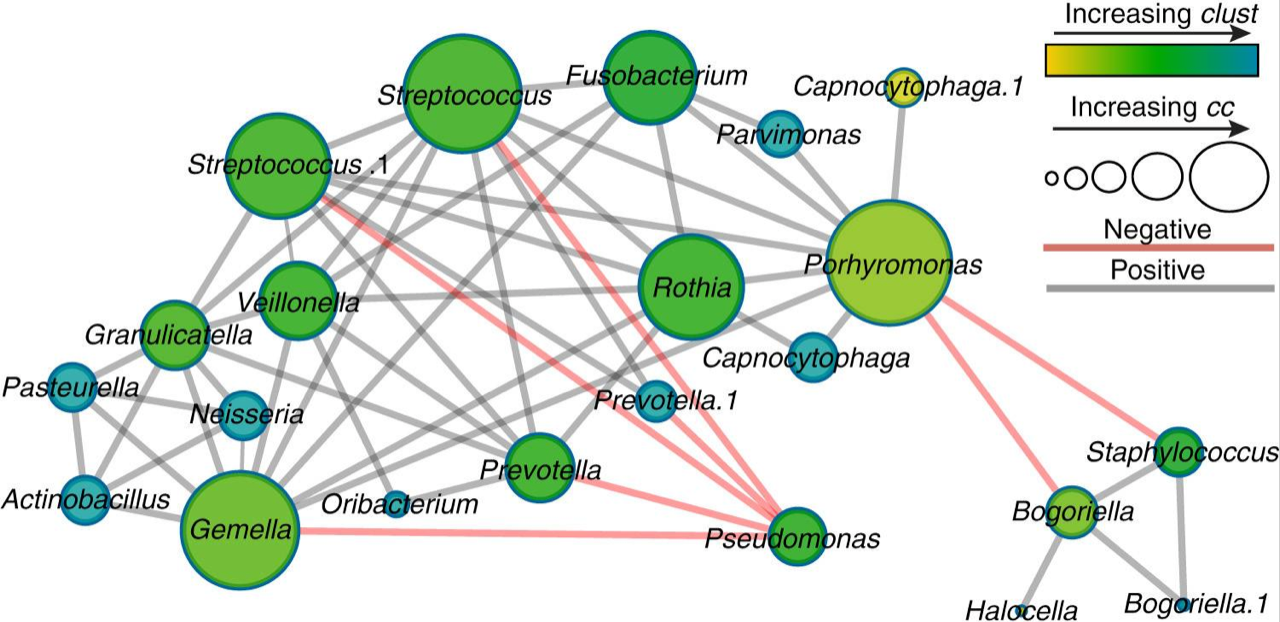
Ecological networking of cystic fibrosis lung infections. Quinn RA, Whiteson K, Lim YW, Zhao J, Conrad D, LiPuma JJ, Rohwer F, Widder S. NPJ Biofilms Microbiomes. 2016 Dec
The origin and evolution of cell types. Arendt D*, Musser JM*, Baker CVH, Bergman A, Cepko C, Erwin DH, Pavlicev M, Schlosser G, Widder S, Laubichler MD, Wagner GP. Nat Rev Genet. 2016 Dec
News and Features:
Science (350/2015): In depth-Using evolution to better identify cell types
Challenges in microbial ecology: building predictive understanding of community function and dynamics. Widder S, Allen R, Pfeiffer T, Curtis TP, Wiuf C, Sloan WT, Cordero OX, Brown SP, Momeni B, Shou W, Kettle H, Flint HJ, Haas AF, Laroche B, Kreft JU, Rainey PB, Freilich S, Schuster S, Milferstedt K, van der Meer JR, Groβkopf T, Huisman J, Free A, Picioreanu C, Quince C, Klapper I, Labarthe S, Smets BF, Wang H, Isaac Newton Institute Fellows, Soyer OS. ISME J. 2016 Nov
Biogeographic variation in the microbiome of the ecologically important sponge, Carteriospongia foliascens. Luter HM, Widder S, Botté ES, Abdul Wahab M, Whalan S, Moitinho-Silva L, Thomas T, Webster NS. PeerJ. 2015 Dec
Wiring for Independence: Positive Feedback Motifs Facilitate Individuation of Traits in Development and Evolution. Pavlicev M, Widder S. J Exp Zool B Mol Dev Evol. 2015 Mar
Fluvial network organization imprints on microbial co-occurrence networks. Widder S, Besemer K, Singer GA, Ceola S, Bertuzzo E, Quince C, Sloan WT, Rinaldo A, Battin TJ. Proc Natl Acad Sci U S A. 2014 Sep
News and Features:
PNAS (111/2014): This week in PNAS
Metagenomic analysis reveals presence of Treponema denticola in a tissue biopsy of the Iceman. Maixner F*, Thomma A*, Cipollini G, Widder S, Rattei T, Zink A. PLoS One. 2014 Jun
News and Features:
Der Kurier, Printversion (16/7/2014): Schon Ötzi wurde von Zahnfleischentzündung gequält.
Der Standard (16/7/2014): Ötzis Knochenanalyse enthüllt "nichtmenschliche" DNA
Universität Wien - uni:view magazin (15/7/2014): Ötzis "nichtmenschliche" DNA entschlüsselt
Deciphering microbial interactions and detecting keystone species with co-occurrence networks. Berry D, Widder S. Front Microbiol. 2014 May
Ultra deep sequencing of Listeria monocytogenes sRNA transcriptome revealed new antisense RNAs. Behrens S, Widder S, Mannala GK, Qing X, Madhugiri R, Kefer N, Abu Mraheil M, Rattei T, Hain T. PLoS One. 2014 Feb
Evolvability of feed-forward loop architecture biases its abundance in transcription networks. Widder S, Solé R, Macía J. BMC Syst Biol. 2012 Jan
Specialized or flexible feed-forward loop motifs: a question of topology. Macía J*, Widder S*, Solé R. BMC Syst Biol. 2009 Aug
Why are cellular switches Boolean? General conditions for multistable genetic circuits. Macía J, Widder S, Solé R. J Theor Biol. 2009 Nov
Monomeric bistability and the role of autoloops in gene regulation. Widder S*, Macía J*, Solé R. PLoS One. 2009
Dynamic patterns of gene regulation I: simple two-gene systems. Widder S, Schicho J, Schuster P. Front Microbiol. 2014 May
A minimal and self-consistent in silico cell model based on macromolecular interactions. Flamm C, Endler L, Müller S, Widder S, Schuster P. Philos Trans R Soc Lond B Biol Sci. 2007 Oct
The SBML ODE Solver Library: a native API for symbolic and fast numerical analysis of reaction networks. Machné R, Finney A, Müller S, Lu J, Widder S, Flamm C. Bioinformatics. 2006 Jun
A generalized model of the repressilator. Müller S, Hofbauer J, Endler L, Flamm C, Widder S, Schuster P. Journal Math Biol. 2006 Dec
p59OASL, a 24–54 oligoadenylate synthetase like protein: a novel human gene related to the 24–54 oligoadenylate synthetase family. Hartmann R, Olsen HS, Widder S, Jørgensen R, Justesen J. Nucleic Acids Research, 1998